AlignCell
Single-cell RNA-seq from raw counts to biological insights
The Challenge
Single-cell RNA-seq reveals cellular heterogeneity invisible to bulk methods, but the analytical complexity scales accordingly. Researchers face decisions at every step: filtering thresholds, normalization approaches, clustering resolution, cell type assignment. Each choice affects downstream conclusions, and the field’s best practices continue to evolve rapidly.
Without specialized expertise, it’s easy to over-cluster noise into spurious populations, miss rare cell types, or misinterpret batch effects as biological variation. The gap between raw 10X output and publishable biological insights requires both technical skill and domain knowledge.
How AlignCell Helps
AlignCell provides a complete single-cell analysis workflow built on Seurat 5.0, the most widely adopted framework in the field. Our 11-step analytical pipeline takes your data from raw counts through quality control, normalization, dimensionality reduction, and clustering to biological interpretation.
Cell type annotation combines automated methods with expert review. We apply reference-based annotation approaches and validate assignments against known marker genes, ensuring cell type labels reflect actual biology rather than algorithmic artifacts.
For multi-sample experiments, Harmony batch correction integrates data across conditions while preserving biological variation. This enables direct comparison of cell populations across treatment groups, timepoints, or patient samples without confounding by technical batch.
When your research questions involve developmental processes or cell state transitions, AlignCell includes trajectory analysis using Slingshot or Monocle3. We identify differentiation paths, pseudotime orderings, and genes that drive cell fate decisions.
Cell-cell communication analysis via CellChat reveals how cell populations interact through ligand-receptor signaling. This is particularly valuable for understanding tissue microenvironments, immune responses, and tumor-stroma interactions.
The AlignCell Interface
11-step guided workflow with Seurat 5.0 integration and dark theme UI:

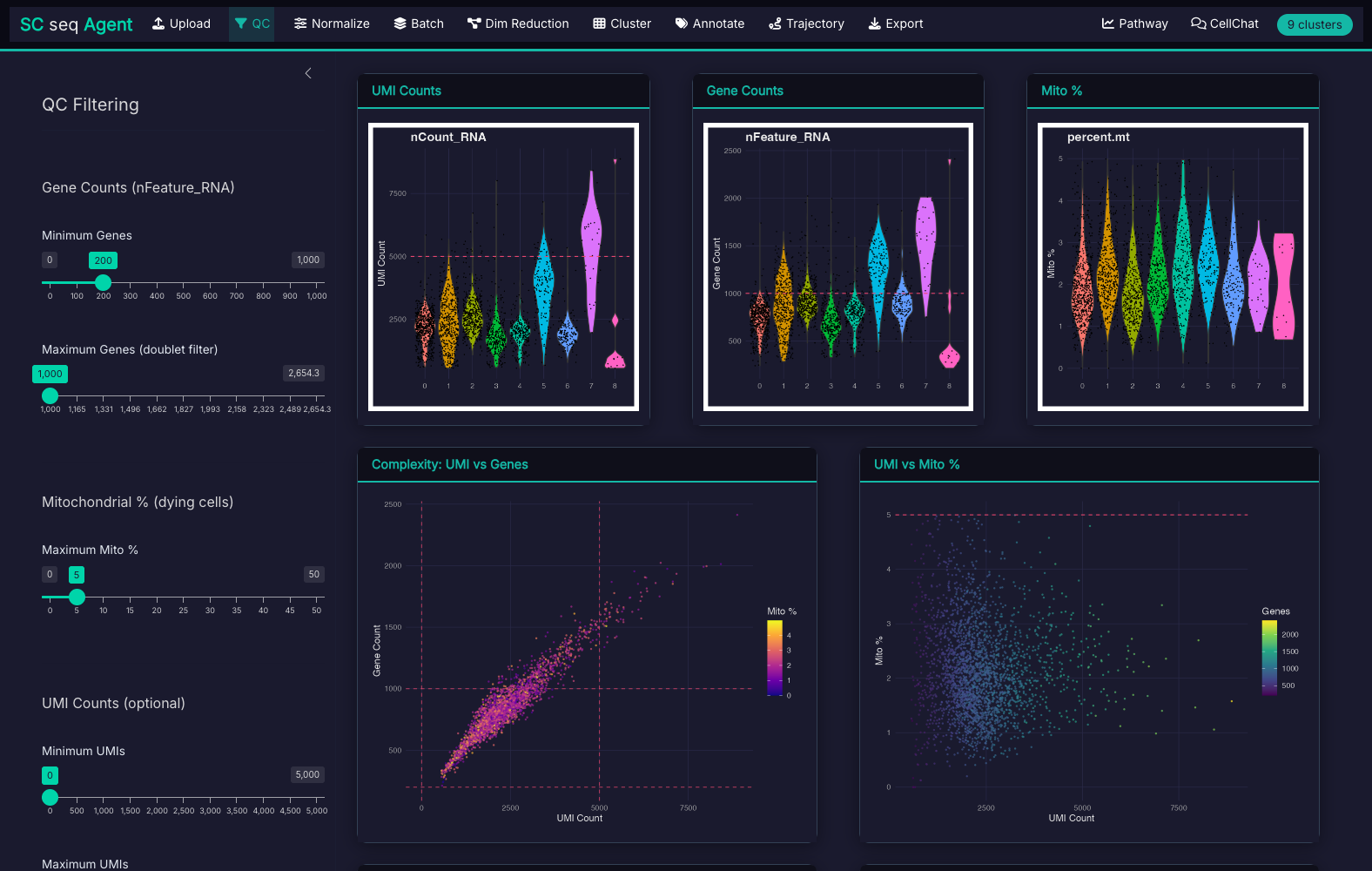


What You Receive
Annotated Seurat Object
Complete Seurat object with cell type annotations, cluster assignments, and all computed embeddings ready for further exploration.
Publication Figures
UMAP visualizations, marker heatmaps, dot plots, and trajectory diagrams formatted for journal submission.
Differential Expression
Marker genes for each cluster, differential expression between conditions, and cell type composition comparisons.
Communication Analysis
CellChat results with ligand-receptor interactions, signaling pathway networks, and cell-cell communication visualizations.
Methodology
Harmony
Slingshot
CellChat
SingleR
SCTransform
AlignCell implements the Seurat 5.0 workflow as documented by the Satija Lab, incorporating field-standard tools including Harmony for batch correction, SingleR for reference-based annotation, and CellChat for communication analysis.
Our 11-step workflow covers: data loading, QC filtering, normalization (LogNormalize or SCTransform), batch correction, PCA, UMAP, clustering, marker detection, cell type annotation, trajectory inference, and pathway/communication analysis.
Ideal For
- 10X Genomics single-cell experiments (any chemistry version)
- Multi-sample studies requiring batch integration
- Developmental or differentiation studies needing trajectory analysis
- Tumor microenvironment characterization
- Immune cell profiling at single-cell resolution
- Any scRNA-seq dataset where cell type composition and communication matter
Start Your Analysis
Ready to analyze your data with AlignCell? Submit your project and we'll scope a plan tailored to your experimental design.