Aligncellscape
General-purpose cell type deconvolution for tumor and PBMC analysis
The Challenge
Cell type deconvolution from bulk RNA-seq is essential for understanding tissue composition, but existing tools often specialize in narrow domains. Researchers studying tumors need stromal signatures that PBMC-focused tools lack. Those analyzing immune cells need fine-grained subtypes that tumor-focused methods miss. Switching between tools creates inconsistency and requires learning multiple interfaces.
The field needs a unified deconvolution platform that handles diverse tissue contexts with validated signatures, combining the depth of specialized tools with the convenience of a single, well-designed interface.
How Aligncellscape Helps
Aligncellscape provides general-purpose cell type deconvolution through three built-in signature modules that cover the most common analysis scenarios. The tumor_microenv module resolves 24 cell types including fibroblasts, CAFs, pericytes, and immune infiltrates. The pbmc_immune module provides 28 fine-grained PBMC cell types derived from the Monaco reference. The hallmark module scores 50 MSigDB pathways for functional interpretation.
The underlying methodology uses robust linear regression (RLM) with validated signatures. This approach was benchmarked on the DREAM Tumor Deconvolution Challenge (mean Pearson r = 0.688-0.727) and independently validated on Wu et al. breast cancer pseudo-bulk data, achieving correlations of 0.82-0.92 for key cell types not present in the DREAM benchmark.
Two analysis modes accommodate different research questions. The deconvolve() function returns compositional proportions that sum to 1.0 per sample, ideal when you need to compare cell type fractions across conditions. The enrich() function returns independent enrichment scores (xCell-style), robust when some cell types are absent from your samples.
The modular signature system is extensible. Beyond the three built-in modules, you can load custom GMT files or create signature modules tailored to your specific tissue context.
The Aligncellscape Interface
Dark theme UI following the AlignMatrix design system:
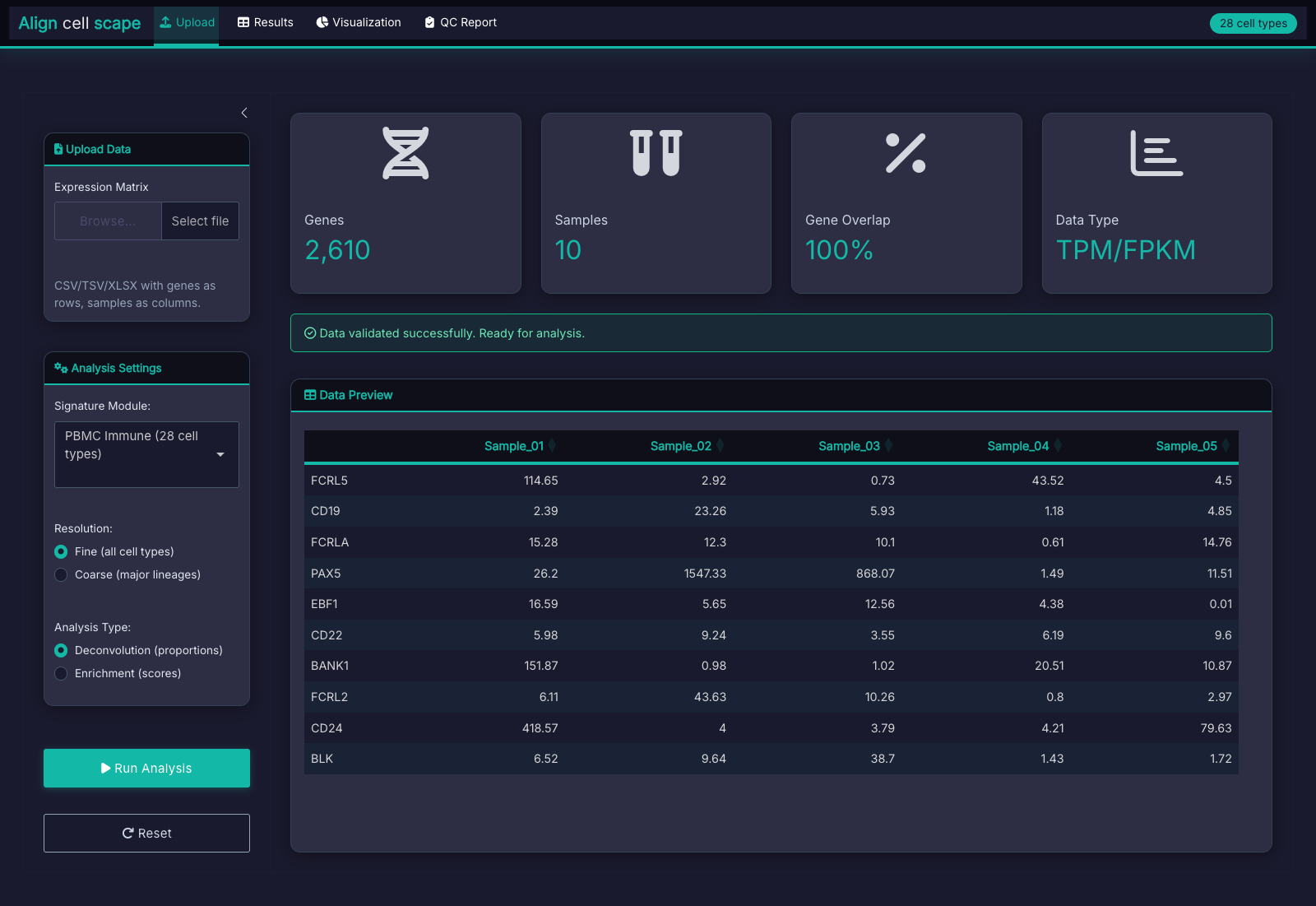
What You Receive
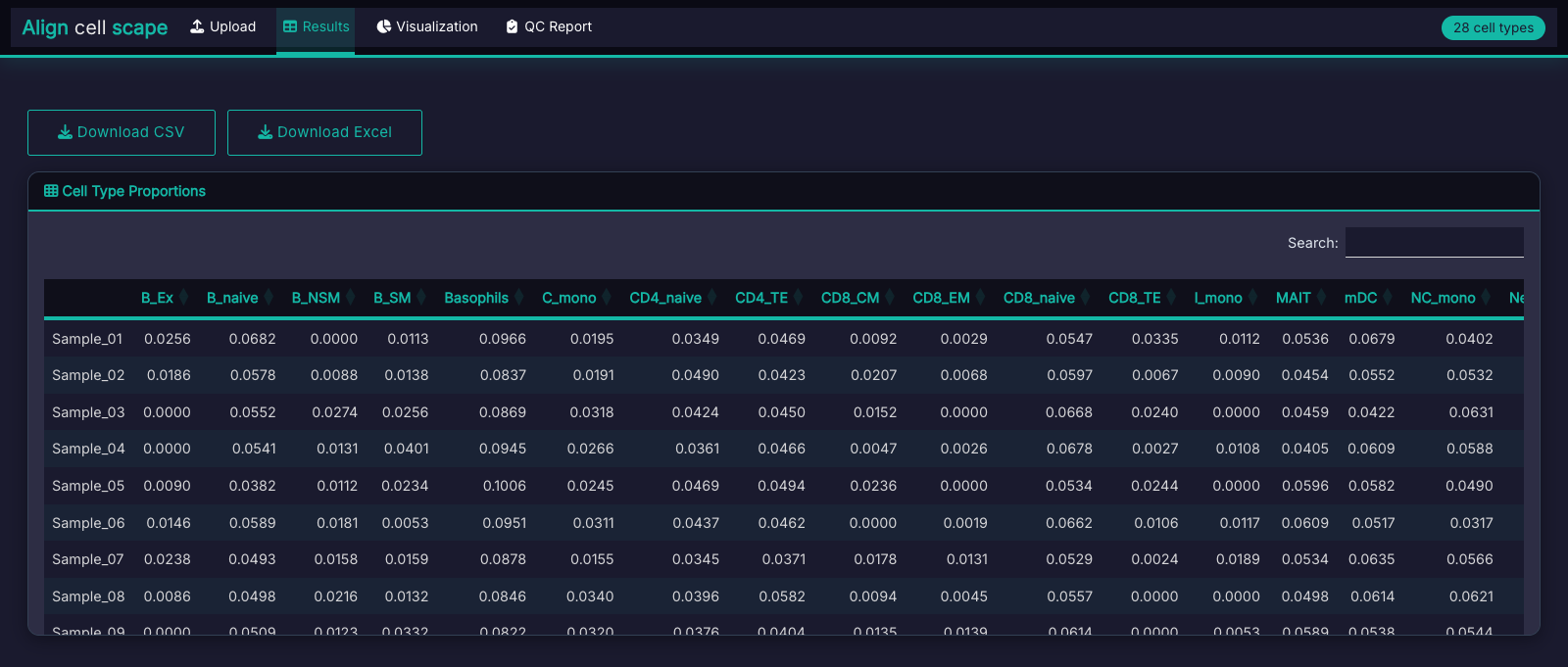
Cell Type Proportions
Estimated proportions for all cell types in your chosen module, with compositional or enrichment-based quantification.
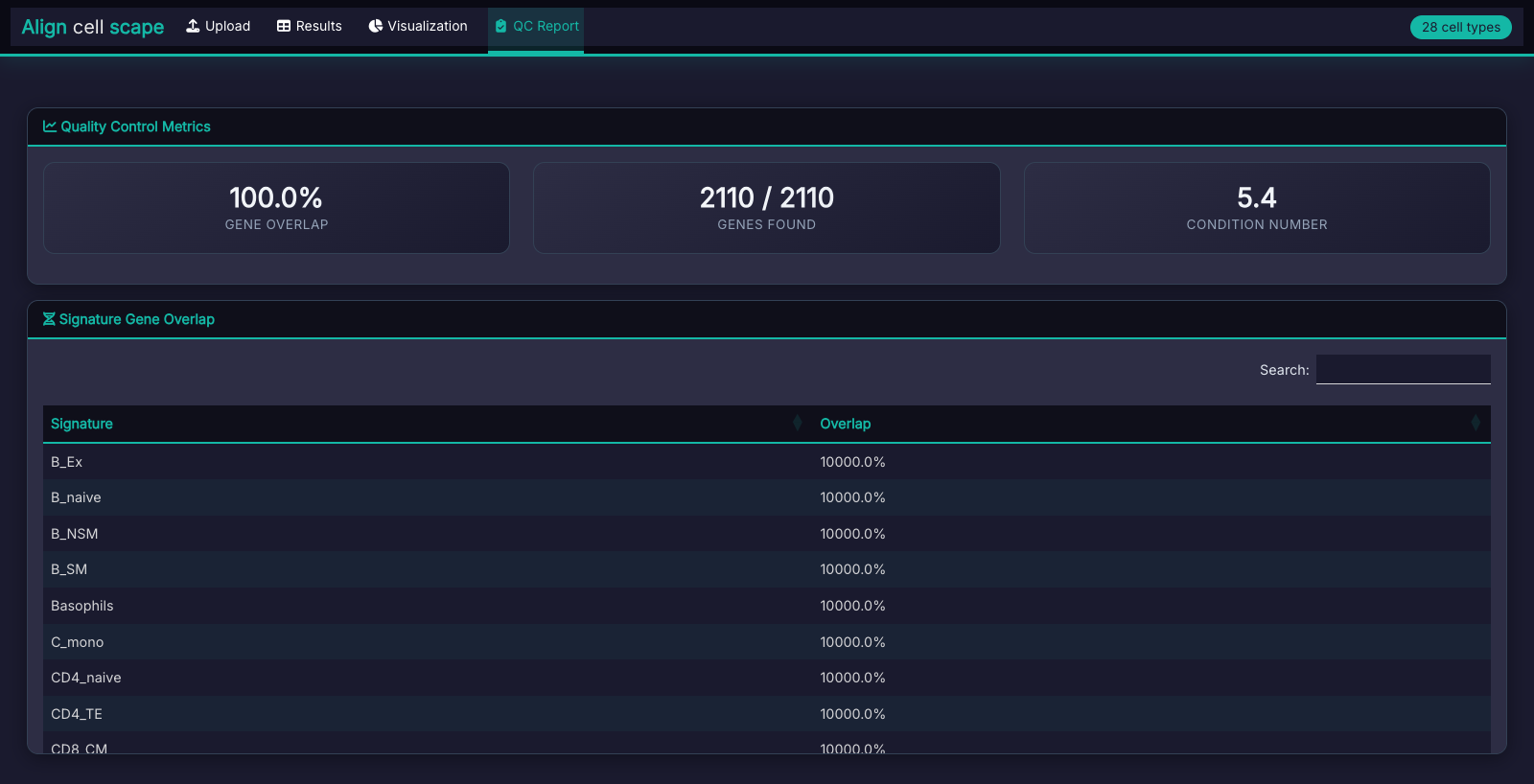
QC Report
Gene overlap statistics, condition number assessment, and warnings for potential deconvolution issues.
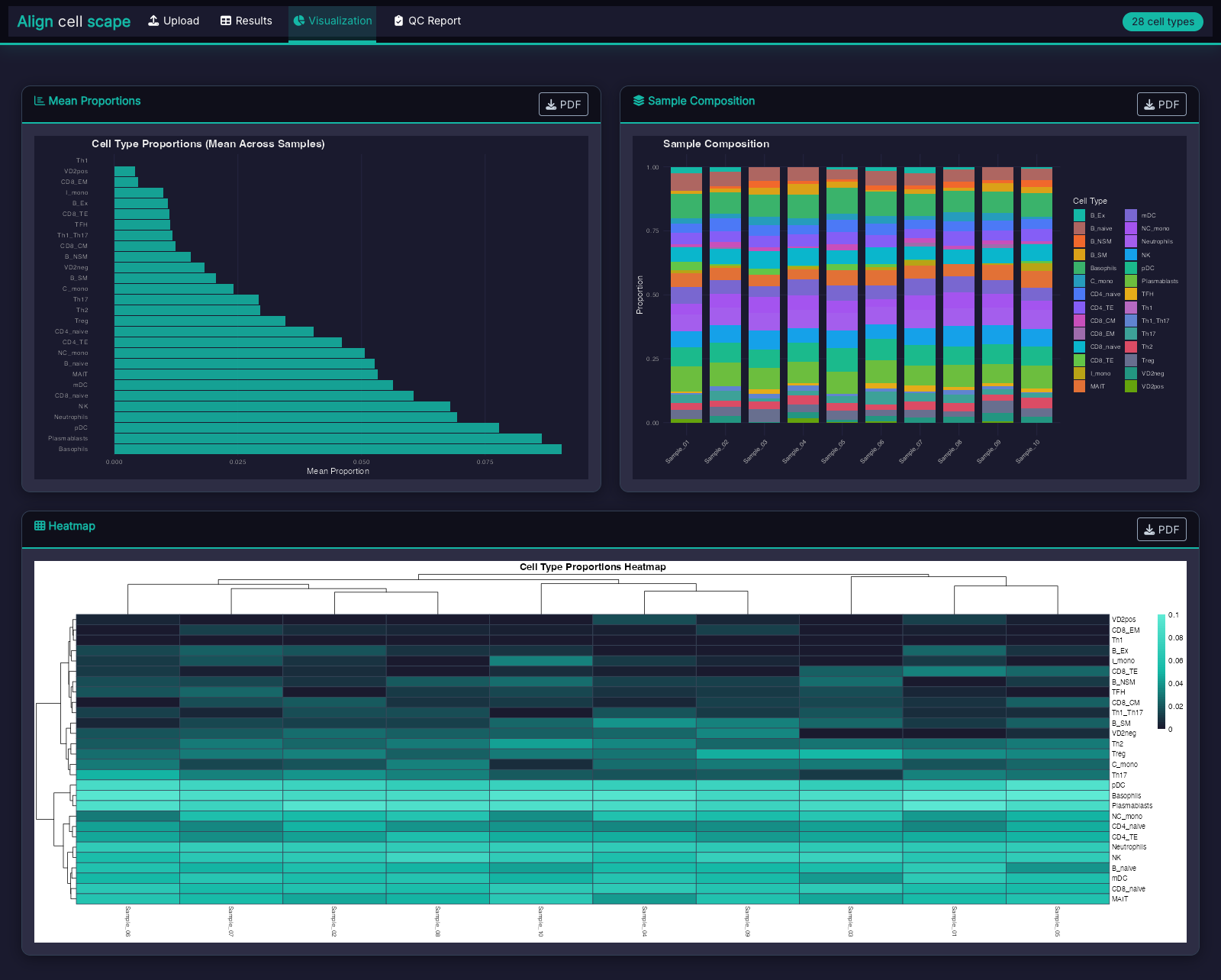
Publication Figures
Dark-themed bar charts, stacked composition plots, and heatmaps formatted for journal submission.
Export Options
CSV tables, PDF figures, and full R objects for downstream analysis in your own pipelines.
Methodology & Validation
xCell-style enrichment
Monaco reference
DREAM validated
3 signature modules
Aligncellscape extends the atlas29 methodology with support for multiple tissue contexts. Signatures derive from curated sources: Monaco et al. (2019) Cell Reports for PBMC immune types, and literature-curated markers for tumor microenvironment populations.
Validation: DREAM Tumor Deconvolution Challenge (mean r = 0.688-0.727); Wu et al. breast cancer pseudo-bulk (r = 0.82-0.92 for pericytes, T-cells, endothelial, cancer epithelial); fibroblast signatures validated at r = 0.811.
Ideal For
- Tumor microenvironment characterization from bulk RNA-seq
- PBMC immune cell profiling with fine-grained subtypes
- Studies requiring both stromal and immune cell quantification
- Fibroblast and CAF analysis in fibrosis or cancer contexts
- Pathway activity scoring alongside cell type deconvolution
- Any bulk RNA-seq where multiple tissue contexts need consistent analysis
Start Your Analysis
Ready to analyze your data with Aligncellscape? Submit your project and we'll scope a plan tailored to your experimental design.